
GutenOCR: A Grounded Vision-Language Front-End for Documents
GutenOCR is a family of vision-language models designed to serve as a ‘grounded OCR front-end’, providing high-quality text transcription and explicit geometric grounding.
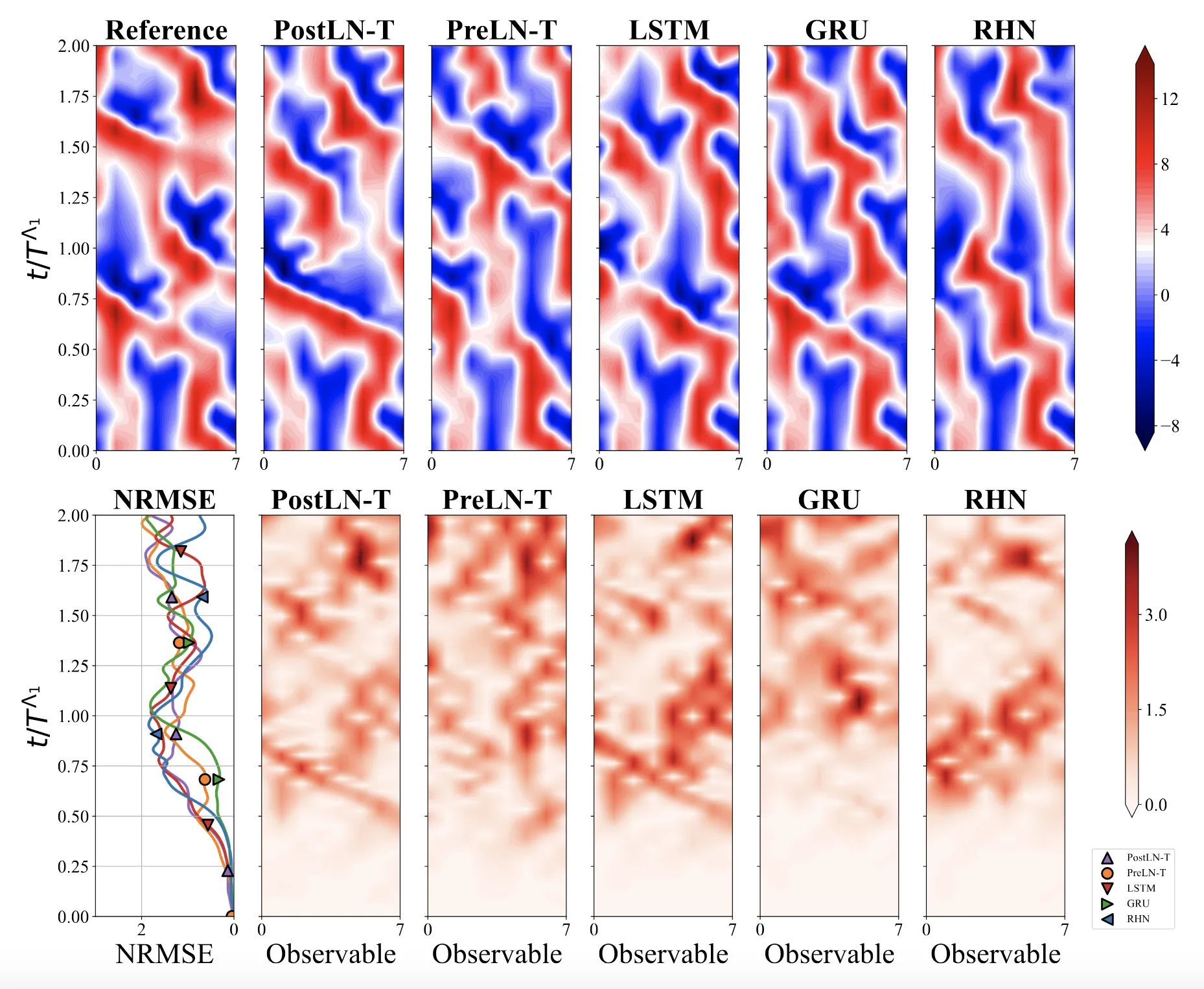
Optimizing Sequence Models for Dynamical Systems
We systematically ablate core mechanisms of Transformers and RNNs, finding that attention-augmented Recurrent Highway Networks outperform standard Transformers on forecasting high-dimensional chaotic systems.
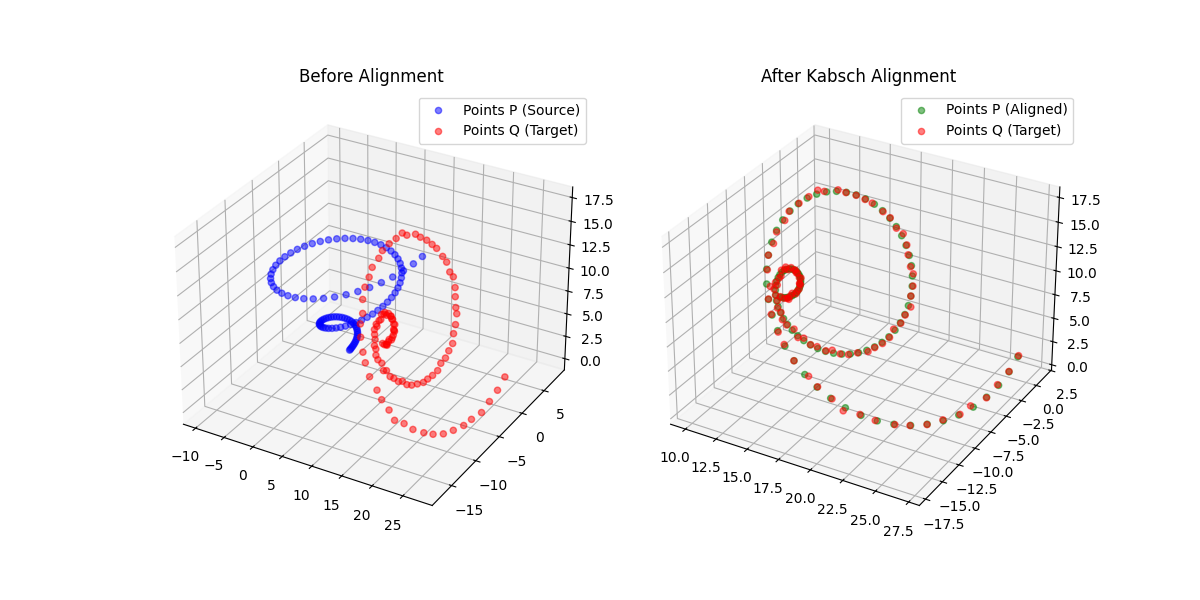
Kabsch-Horn Cookbook: Differentiable Alignment
A differentiable point-set alignment library implementing N-dimensional Kabsch, Horn quaternion, and Umeyama scaling algorithms with per-point weights, batch dimensions, and custom autograd across NumPy, PyTorch, JAX, TensorFlow, and MLX.
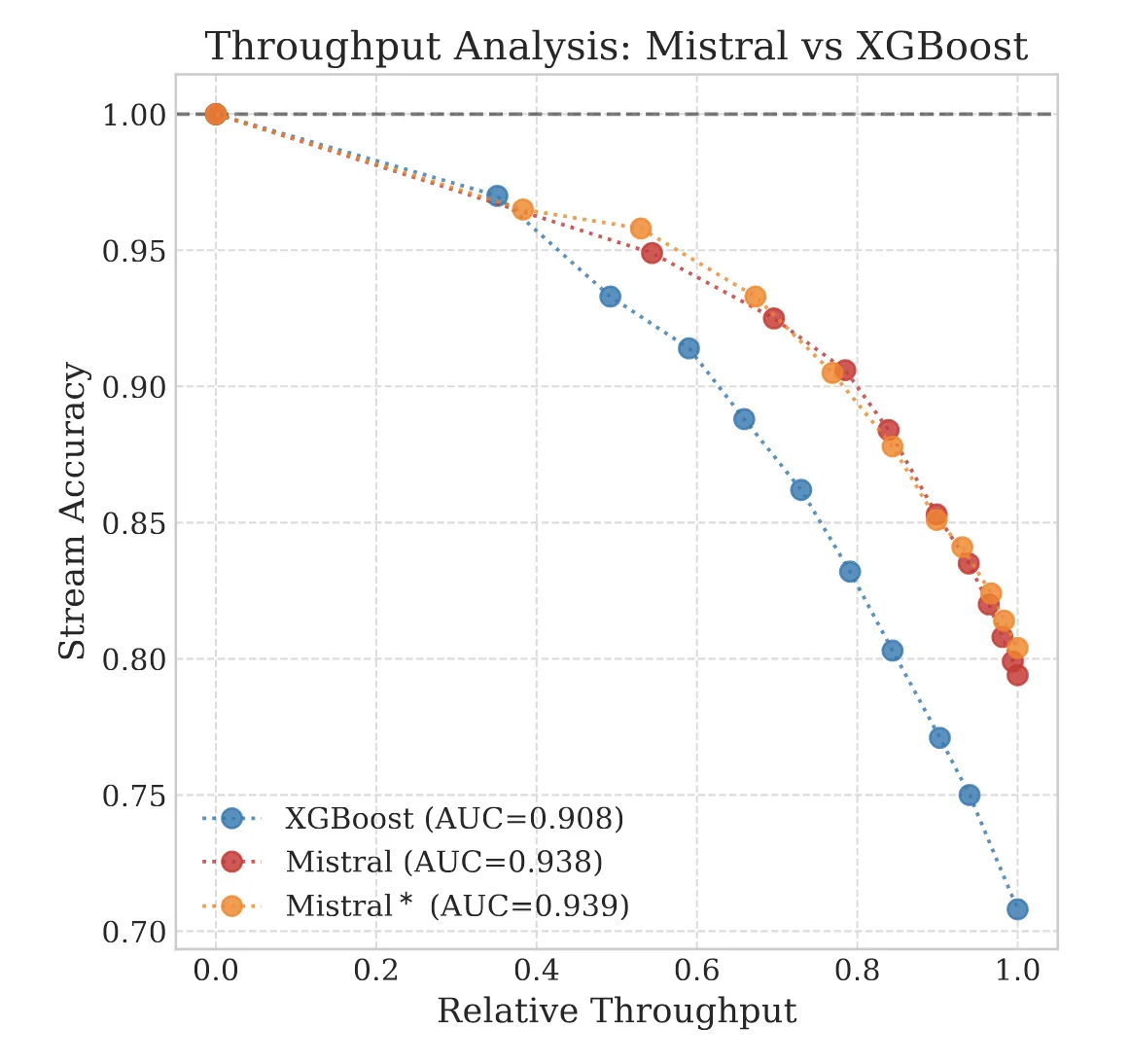
The Reliability Trap: The Limits of 99% Accuracy
We explore the ‘Silent Failure’ mode of LLMs in production: the limits of 99% accuracy for reliability, how confidence decays in long documents, and why standard calibration techniques struggle to fix it.
The Evolution of Page Stream Segmentation: Rules to LLMs
We trace the history of Page Stream Segmentation (PSS) through three eras (Heuristic, Encoder, and Decoder) and explain how privacy-preserving, localized LLMs enable true semantic processing.
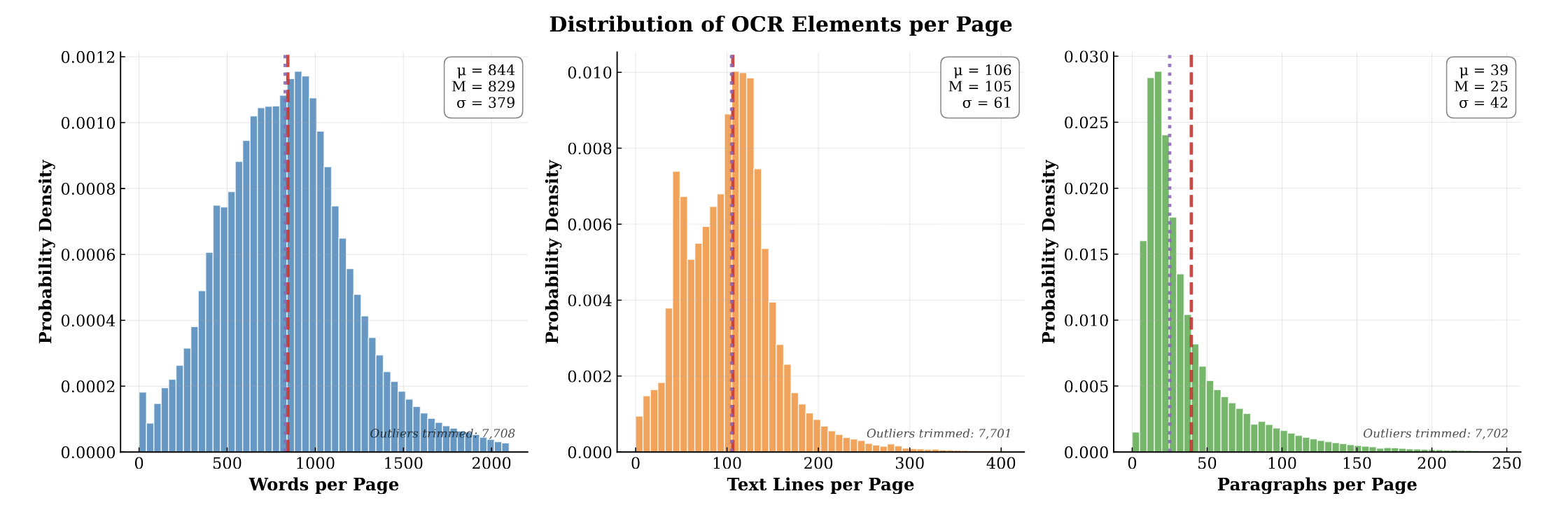
PubMed-OCR: PMC Open Access OCR Annotations
PubMed-OCR provides 1.5M pages of scientific articles with comprehensive OCR annotations and bounding boxes to support layout-aware modeling and document analysis.
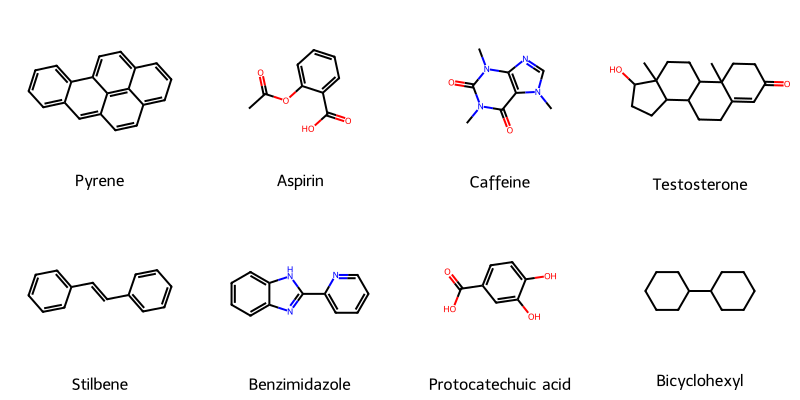
Molecular String Renderer: Chemical Visualization Library
An RDKit wrapper treating molecular visualization as a software engineering problem, implementing strategy pattern for SVG generation with automatic raster fallback, native SELFIES support for generative AI workflows, and strict type safety for batch processing in molecular ML training pipelines.
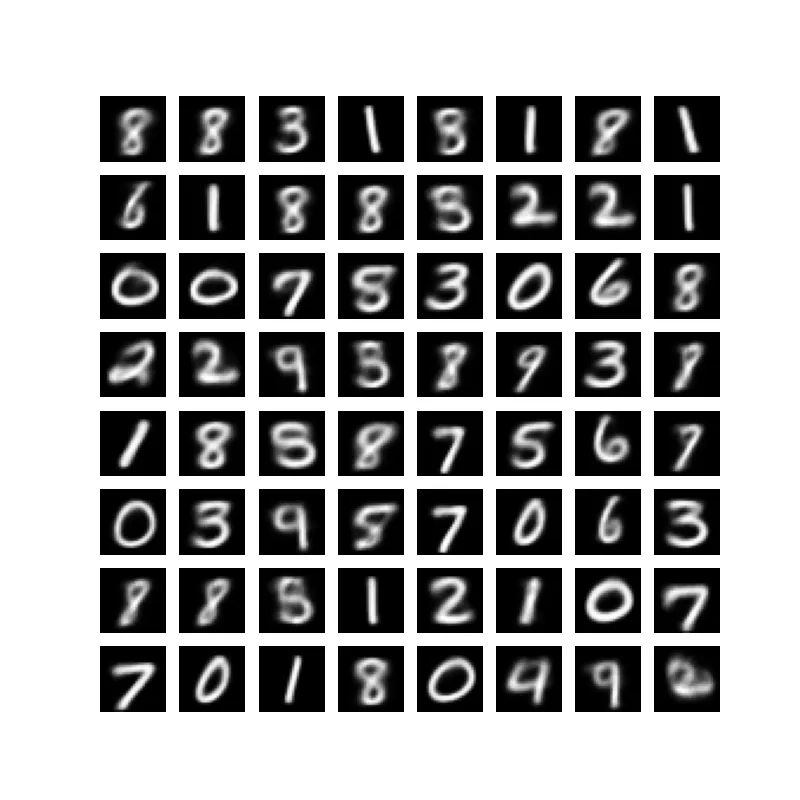

Importance Weighted Autoencoders: Beyond the Standard VAE
Discover how Importance Weighted Autoencoders (IWAEs) use the same architecture as VAEs with a different objective that optimizes a tighter bound on the log-likelihood, leveraging multiple samples effectively.
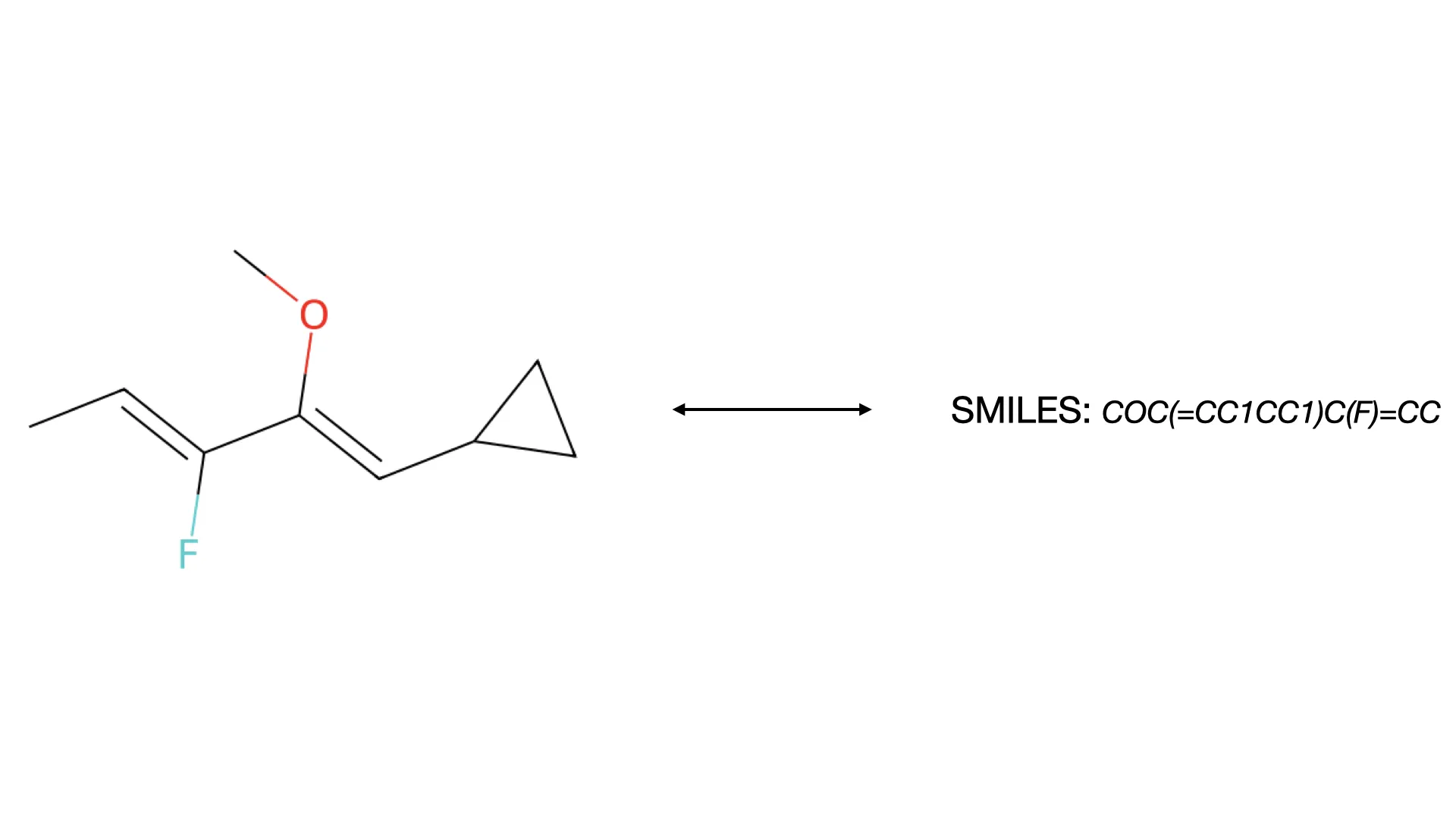
What is Optical Chemical Structure Recognition (OCSR)?
Discover how OCSR technology bridges the gap between molecular images and machine-readable data, evolving from rule-based systems to modern deep learning models for chemical knowledge extraction.
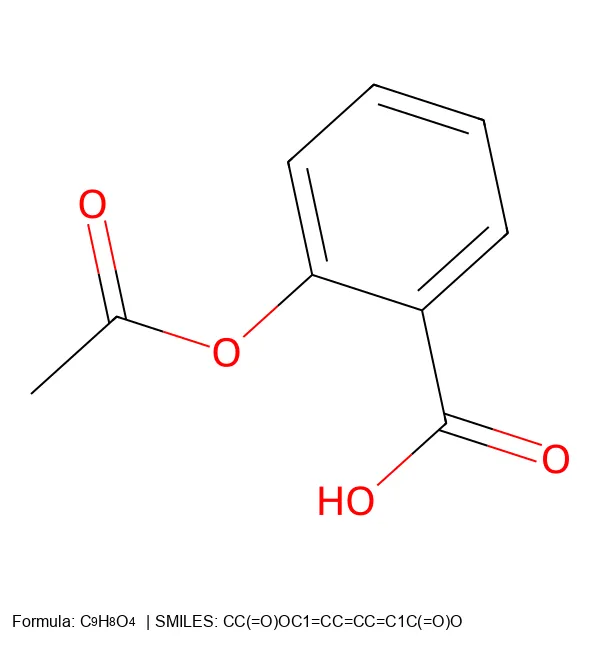
Converting SMILES and SELFIES to 2D Molecular Images
Build a Python CLI tool that converts SMILES and SELFIES notation into 2D molecular images with chemical formulas and legends, including an SVG path for figures.

Exponential Random Numbers: Two Classic Algorithms
Explore two fundamental approaches to generating exponentially distributed random numbers: the modern inverse transform method using logarithms and von Neumann’s ingenious 1951 comparison-based algorithm that avoids transcendental functions entirely.
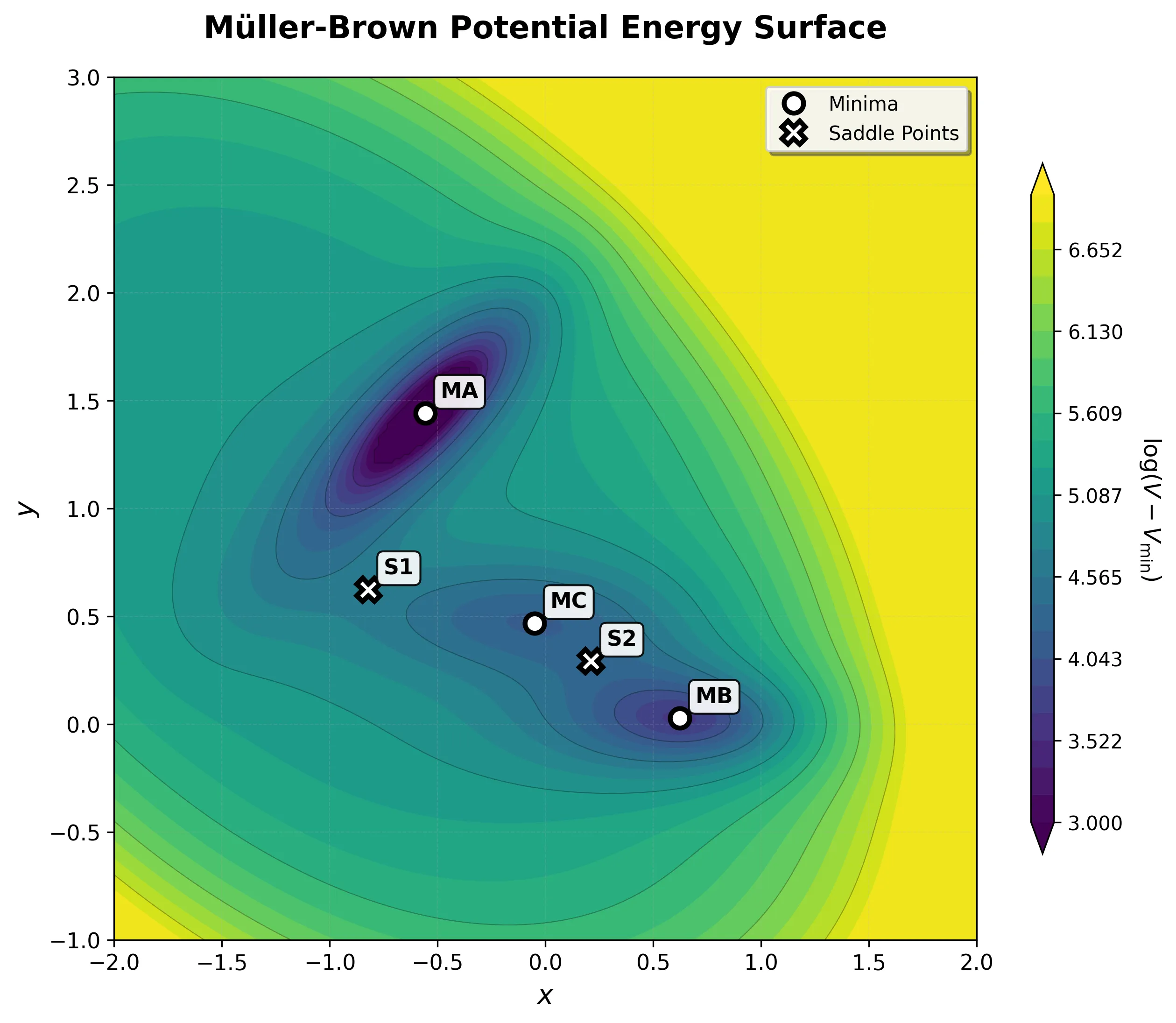
Implementing the Müller-Brown Potential in PyTorch
Step-by-step implementation of the classic Müller-Brown potential in PyTorch, with performance comparisons between analytical and automatic differentiation approaches for molecular dynamics and machine learning applications.