Abstract
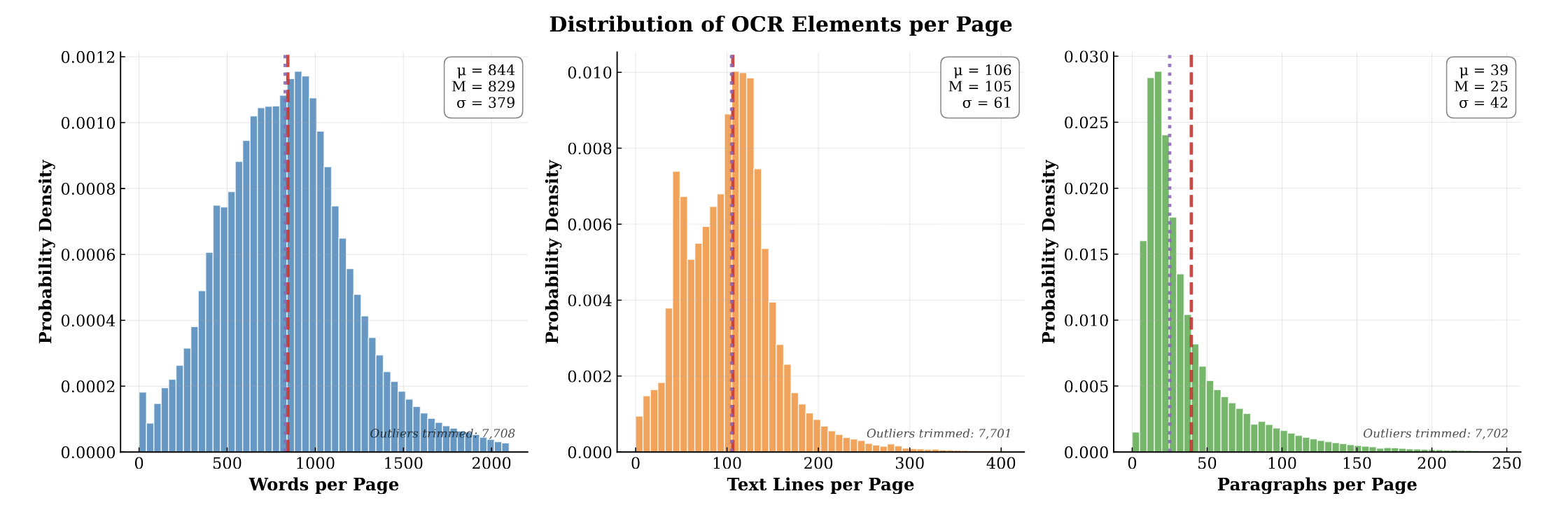
PubMed-OCR is an OCR-centric corpus of scientific articles derived from PubMed Central Open Access PDFs. Each page image is annotated with Google Cloud Vision and released in a compact JSON schema with word-, line-, and paragraph-level bounding boxes. The corpus spans 209.5K articles (1.5M pages; ~1.3B words) and supports layout-aware modeling, coordinate-grounded QA, and evaluation of OCR-dependent pipelines. We analyze corpus characteristics (e.g., journal coverage and detected layout features) and discuss limitations, including reliance on a single OCR engine and heuristic line reconstruction. We release the data and schema to facilitate downstream research and invite extensions.
Key Contributions
- OCR-First Supervision: Unlike prior datasets for PubMed that align XML to PDFs, PubMed-OCR provides native OCR annotations (Google Cloud Vision), bypassing alignment errors and covering non-digital scanned pages.
- High-Density Annotation: At ~1.3B words across 1.5M pages, PubMed-OCR delivers 10x higher word density and 4x finer line granularity than comparable industrial OCR corpora like OCR-IDL.
- Multi-Level Bounding Boxes: Includes explicit word-, line-, and paragraph-level bounding boxes to support hierarchical document understanding and layout-aware modeling. We also hope that this leads to VQA datasets with grounded answers in document layout.
- Open Access & Reproducibility: Derived strictly from the redistributable PMCOA subset, releasing both the JSON annotations and original PDFs to ensure verifiable and reproducible research.
Technical Implementation
Corpus Construction
PubMed-OCR is built from PubMed Central Open Access (PMCOA) PDFs, chosen specifically because the PMCOA license permits redistribution of both the original documents and derived annotations. Each PDF is rendered to page images, then passed to the Google Cloud Vision (GCV) API. Each page produces a structured JSON annotation file capturing the detected text along with bounding box geometry at word, line, and paragraph levels.
JSON Annotation Schema
Each page annotation follows this compact schema. Bounding boxes are axis-aligned rectangles in [x1, y1, x2, y2] pixel coordinates. Words, lines, and paragraphs are stored as parallel flat lists under the text key:
{
"text": {
"words": [
{"text": "Example", "box": [180, 746, 210, 786]}
],
"lines": [
{"text": "Example sentence", "box": [180, 746, 540, 786]}
],
"paragraphs": [
{"text": "Example sentence\nSecond line", "box": [180, 746, 540, 820]}
]
},
"image": "..."
}

Line Reconstruction
GCV returns word-level detections natively. Line and paragraph groupings are reconstructed using spatial heuristics: words are clustered into lines by vertical overlap and horizontal proximity, and paragraph grouping follows a similar process at a coarser scale. These heuristics work well for standard single-column scientific layouts but can fail on multi-column or irregularly structured pages (see Limitations).
Using the Dataset
The corpus spans 1.5M pages, so streaming is recommended for most use cases:
import json
from datasets import load_dataset
# Streaming is recommended for the full 1.5M-page corpus
ds = load_dataset("rootsautomation/pubmed-ocr", streaming=True, split="train")
# Inspect a page
page = next(iter(ds))
print(f"Article: {page['accession_id']}, Page: {page['page']}")
# Parse OCR annotations
ocr = json.loads(page["ocr_json"])
text = ocr["text"]
# Iterate over lines and words
for line in text["lines"][:5]:
print(f" Line: {line['text']}")
print(f" BBox: {line['box']}")
# Access individual word detections
for word in text["words"][:5]:
print(f" Word: {word['text']}, BBox: {word['box']}")
Full schema documentation is available on the HuggingFace dataset card.
Why This Matters
The lack of large-scale, high-quality OCR datasets with explicit geometric grounding has been a major bottleneck for training layout-aware models. By releasing PubMed-OCR, we provide the community with the dense, multi-level bounding box annotations necessary to build the next generation of document understanding systems. This dataset directly supports the development of models like GutenOCR, enabling them to learn precise token-to-pixel alignment and robust layout reasoning.
Limitations
- Single OCR engine: All annotations come from Google Cloud Vision. GCV’s error modes (handwriting, degraded scans, complex math, non-Latin scripts) propagate uncorrected into the dataset. Different OCR engines could yield different coverage patterns and error distributions.
- Heuristic line reconstruction: Spatial word-to-line clustering is approximate. Multi-column layouts, rotated text, or unusual page orientations may produce incorrect line groupings.
- PMCOA scope: Coverage is limited to the Open Access subset of PubMed Central. Commercial or subscription articles are excluded.
Citation
@misc{heidenreich2026pubmedocrpmcopenaccess,
title={PubMed-OCR: PMC Open Access OCR Annotations},
author={Hunter Heidenreich and Yosheb Getachew and Olivia Dinica and Ben Elliott},
year={2026},
eprint={2601.11425},
archivePrefix={arXiv},
primaryClass={cs.CV},
url={https://arxiv.org/abs/2601.11425},
}
Related Work
This dataset directly enables GutenOCR, a family of vision-language models trained on PubMed-OCR annotations to produce grounded OCR outputs with explicit bounding boxes.
For related work on document processing pipelines that consume OCR output, see LLMs for Page Stream Segmentation and Page Stream Segmentation with LLMs: Challenges and Applications.